The PCRBIO Rapid Extract PCR Kit and Lysis Kit offer rapid DNA extraction in a convenient, easy-to-use format. The kits contains a lysis and protease buffer system that allows the generation of PCR-ready DNA for use in downstream PCR or qPCR reactions. They are particularly suited to solid tissue such as mouse tail or mouse ear, and can also be used with soft tissue, hair follicle, buccal swab, mammalian blood and FFPE tissue.
Features
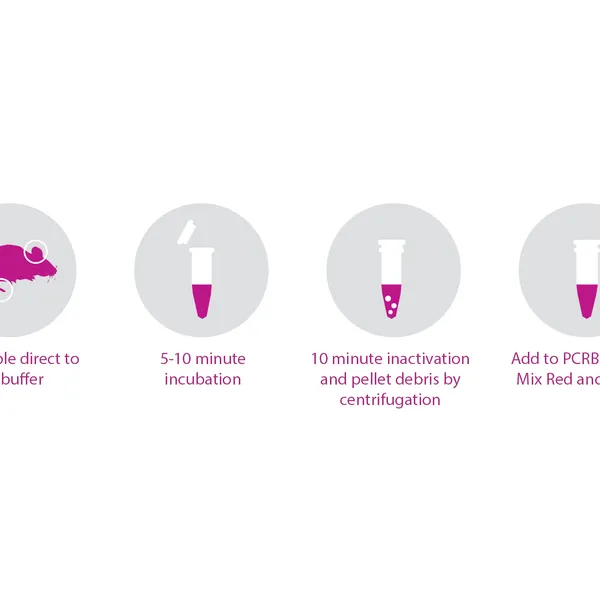
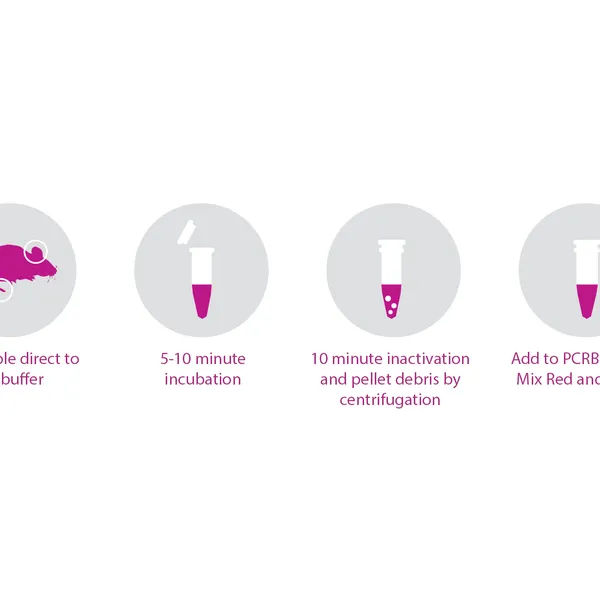
- Rapid, convenient, single-tube DNA extraction
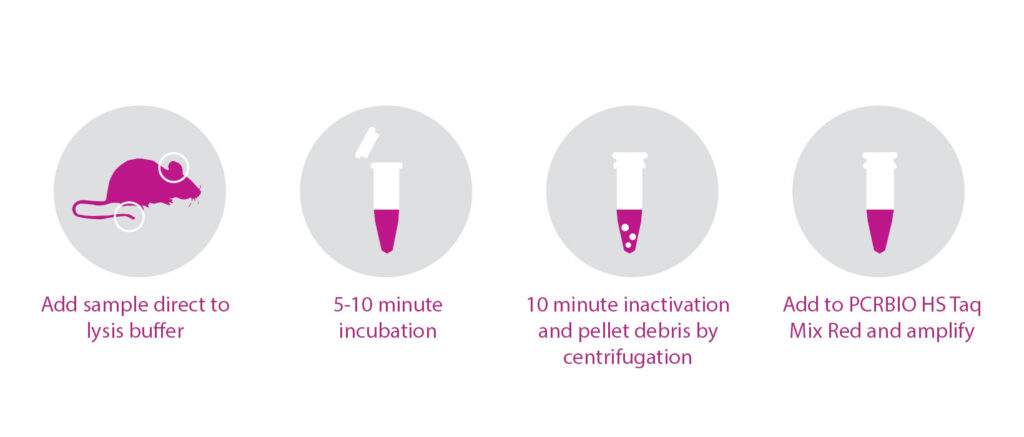
- Produces high yield, PCR-ready DNA in as little as 15 minutes
- Single-tube extraction reduces potential contamination and sample loss
- No need for hazardous chemicals or multiple washing steps
- Available in a PCR kit or lysis-only kit
- PCR kit powered by PCRBIO HS Taq Mix Red for direct gel loading – saving you even more time
- Maximises PCR speed, yield and specificity
- Ideal for complex templates
Sample Types
- Mouse tail clip and ear punch
- Animal tissue
- Hair follicle
- Buccal swab
- Mammalian blood
- FFPE tissue
PCRBIO Rapid Extract PCR Kit
The PCRBIO Rapid Extract PCR Kit combines rapid DNA extraction with fast, highly specific DNA amplification in a single tube. Extracted DNA is amplified in a proprietary buffer system using PCRBIO HS Taq Mix Red.
Our antibody-mediated hot start polymerase uses the latest developments in polymerase technology and buffer chemistry to enhance PCR speed, yield and sensitivity. Primer dimer formation and non-specific amplification are prevented giving superior specificity ideal for complex templates such as mammalian genomic DNA. The mix contains a red dye suitable for direct gel loading without the need for additional loading buffer.
PCRBIOaa Rapid Extract Lysis Kit
PCRBIO Rapid Extract Lysis Kit contains only the lysis and protease buffer system allowing the generation of PCR-ready DNA for use in downstream PCR or qPCR reactions.
Applications
- Genotyping
- Transgene detection
- Knockout analysis
- Sequencinga
PCRBIO Rapid Extract PCR Kit
| Component | 80 Reactions | 400 Reactions |
|
5x PCRBIO Rapid Extract Buffer A |
1 x 1.6mL |
5 x 1.6mL |
|
10x PCRBIO Rapid Extract Buffer B |
1 x 800μL |
5 x 800μL |
|
2x PCRBIO HS Taq Mix Red |
2 x 1mL |
10 x 1mL |
PCRBIO Rapid Extract Lysis Kit
| Component | 80 Extractions | 240 Extractions |
|
5x PCRBIO Rapid Extract Buffer A |
1 x 1.6mL |
3 x 1.6 mL |
|
10x PCRBIO Rapid Extract Buffer B |
1 x 800μL |
3 x 800 μL |
| Reaction Volume | Storage |
|
50μL |
On arrival, products should be stored between -30 and -20 °C. If stored correctly the kit will retain full activity until the indicated expiry date. The kit may also be stored at 4 °C for up to 1 month. |
Product Flyers
Product Manuals
Material Safety Data Sheets
Certificate of Analysis Finder
Frequently Asked Questions (FAQs)
Yes. Here is an easy protocol for DNA extraction from nematodes.
- Prepare the extraction solution by combining 2 μL 5x Rapid Extract Buffer A with 1 μL Rapid Extract Buffer B and 7 μL PCR-grade water (Cat. no. PB40.40).
- Add 1 nematode (these volumes can also work for multiple nematodes per prep).
- Incubate the samples for 5 min at 75 °C for sample lysis.
- Deactivate the reation by incubating for 10 min at 95 °C.
- At 90 μL to the deactivated extraction mix and centrifuge at maximum speed (~13,500 g) in a microcentrifuge for 1 min.
- The supernatant can be used as template in PCR and qPCR immediately, or stored at -20 °C for later use.
- Generally, 1-2 μL of supernatant is sufficient as template in a 20 μL PCR reaction.
FFPE samples should be prepared as follows:
Deparaffinisation:
Remove the paraffin wax from the FFPE tissue section using a solvent like xylene or a specialized buffer.
Xylene is the most widely used solvent in the deparaffinisation of tissue sections. It dissolves the paraffin wax, enabling it to be removed from the tissue. For effective deparaffinization, the slides are typically immersed in xylene or a xylene substitution for 2–3 cycles, with each immersion lasting between 5 and 10 minutes. Xylene is highly effective but also toxic, so appropriate safety measures – such as working in a well-ventilated space or using xylene substitutes – should be followed.
The first immersion into the xylene bath begins dissolving the paraffin. After 5-10 minutes, the slides are transferred to the next bath. A second xylene bath ensures any remaining paraffin is fully dissolved, further improving the quality of deparaffinisation. Researchers may choose to perform a third xylene bath for thicker or older sections.
Rehydration:
After the paraffin is removed, the tissue needs to be rehydrated to restore its original state. This is done by gradually decreasing the concentration of ethanol, which is critical to prevent tissue damage. This step also helps eliminate any residual xylene that may still be present in the sample.
The rehydration process typically follows these 4 steps:
- 100% Ethanol: The slides are immersed in absolute ethanol for 5 minutes to remove any remaining xylene.
- 95% Ethanol: The slides are transferred to a solution of 95% ethanol for another 5 minutes. This step starts the gradual rehydration process.
- 70% Ethanol: The next step involves placing the slides in 70% ethanol for 5 minutes, which further hydrates the tissue.
- 50% Ethanol: The slides are then moved to 50% ethanol for 5 minutes to ensure full hydration.
By the end of this step, the tissue has been completely rehydrated and is ready for further analysis.
Extraction:
Follow our protocol for PCRBIO Rapid Extract products. Typical incubation times could be increased based on the size/type of tissue, up to 48 hours (try few time points to identify optimal conditions when doing this for the first time). Optimal conditions depend on the tissue type and specific workflow.
This non-inhibitory dye, added to enable direct gel loading, runs at a similar rate to 200-300 bp DNA fragments on a 1% agarose gel and at 50-100 bp on a 2% agarose gel. You may notice a shift in this apparent molecular weight when running gels of different agarose content.
PCRBIO Rapid Extract PCR Kit contains 5x PCRBIO Rapid Extract Buffer A (Lysis buffer), 10x PCRBIO Rapid Extract Buffer B (Protease buffer) and 2x PCRBIO HS Taq Mix Red.
PCRBIO Rapid Extract Lysis Kit contains 5x PCRBIO Rapid Extract Buffer A (Lysis buffer) and 10x PCRBIO Rapid Extract Buffer B (Protease buffer).
PCRBIO Rapid Extract PCR Kit has the same contents as PCRBIO Rapid Extract Lysis Kit plus PCRBIO HS Taq Mix Red for subsequent PCR amplification.
Rapid Extract PCR Kit was tested with the following list of tissues
- Mammalian cell cultures
- Animal tissues from a variety of organisms (Fish, lobster, Mice tails / ears, Zebrafish, Nematodes, Kidney, Hair Follicle)
- Insects like Drosophila, mosquitos, beetle (legs, wings)
- Buccal swabs
- Gram positive bacteria
- Fungal Mycelia
- Stool samples
- FFPE
- Blood1
- Fish Scale clips
The following tissues can require a pre-wash step before starting the PCRBIO Rapid Extract protocol2
- Yeast2,3 (pre-wash or digestion with zymolase to obtain spheroblasts)
- Feather4 (requires modification of the protocol see below )
- Stool sample5 (requires dilution series to get rid of inhibitors)
If you have a small amount of starting material, we recommend keeping the extracted DNA in the lowest volume possible and to avoid any further dilutions after inactivating the protease.
- Batubara, A. et al. 2016. Study of BMP15 gene polymorphism in Boer, Kacang, and Boerka goats. Jurnal Ilmu Ternak Dan Veteriner 21(4): 224-230. DOI: http://dx.doi.org/10.14334/jitv.v21i4.1636 (2016).
- Inglis, P.W. et al. 2018. Fast and inexpensive protocols for consistent extraction of high quality DNA and RNA from challenging plant and fungal samples for high throughput SNP genotyping and sequencing Applications. PLOS ONE | https://doi.org/10.1371/journal.pone.0206085 October 18.
- Suzuki, T. & Iwahashi, Y. 2013. RNA preparation of Saccharomyces cerevisiae using the digestion method may give misleading results. Appl Biochem Biotechnol 169:1620–1632 DOI 10.1007/s12010-012-0051-8
- Bello, N. et al. 2001. Isolation of genomic DNA from feathers. J Vet Diagn Invest 13:162–164
- Nantavisai, K. et al. 2007. Evaluation of the sensitivities of DNA extraction and PCR methods for detection of Giardia duodenalisin stool specimens. J. Clinical Microbiology. 45: 581–583 doi:10.1128/JCM.01823-06
Protease is a very active enzyme, and if it isn’t properly deactivated through the heat denaturation step, it could quickly degrade the polymerase during the PCR step.
Changes in the colouration of the supernatant could be due to either pigments being released from the tissue into the supernatant or a change of pH. Changes in the supernatant colouration don’t necessarily affect the PCR reaction.
We recommend storing the extracted DNA at -20°C in the short term but for long term storage, we recommend purifying the DNA and re-suspending it in a standard buffer.
The extracted DNA should be suitable for manufacturer’s PCR kit. A version of the PCRBIO Rapid Extract PCR Kit is available without the PCRBIO HS Taq Mix Red: PCRBIO Rapid Extract Lysis kit (PB15.11).
Yes, PCRBIO Rapid Extract Lysis Kit can be used to create template for qPCR. Use PCRBIO Rapid Extract Lysis Kit to prepare the template, and then use that template in a qPCR reaction as specified by the qPCR kit instructions.
For Formalin-Fixed Paraffin-Embedded (FFPE) tissue specimens, we recommend 1 – 2 mm2 of a 10 µm cross-section. If you use less than that, you may not get enough DNA for PCR.
PCRBIO Rapid Extract PCR Kit can be used with human blood however, blood components may inhibit the PCR reaction. Therefore, it’s a good idea to perform a serial dilution of the extracted DNA in order to find the optimal template concentration for the PCR amplification.
If the amplification is less than optimal, we recommend performing an isopropanol/acetate precipitation of the DNA prior to the PCR reaction.
Yes, it can be used to amplify DNA from bacterial colonies or bacterial cell lysate however, it’s not necessary to use the Rapid Extract PCR Kit in order to perform routine colony PCR. If you’re working from bacterial colonies use a sterile tip to pick a colony and resuspend into the 50µl PCR reaction. If working from liquid culture add 5µl of overnight culture to the final mix. Follow the general protocol and increase the initial denaturation time to 10 min at 95°C.
For qPCR, it may be important to extract DNA with the PCRBIO Rapid Extract Lysis Kit to decrease background noise and control the amount of DNA added to the reaction.
Our PCR mixes work reliably with 10 or more copies of template per reaction but not lower than that. Keep the extracted DNA in the lowest volume possible and don’t dilute further after inactivating the protease. We advise setting up the assay with more cells and once the conditions are established, scaling it down from there.
200 mM DTT should be added to the feather and left to incubate overnight in order to break up the keratin.
The PCRBIO HS Taq Mix Red contains a red dye for loading as well as tracking DNA during electrophoresis. In a 2% agarose TAE gel the dye migrates at a rate equivalent to DNA the size of 350bp whilst in a 1% agarose TAE gel the dye migration rate is equivalent to DNA the size of 600bp.






Read more